Saving lives, money and reputation with a fast, new method of bacterial testing in food
Genome guided MALDI TOF offers clear identification of Bacillus-cereus-group dangers
Valentin Pflüger, Mabritec AG
A massive recall of tainted baby formula took place in January 2026. The formula had been tested according to current food safety regulations and found to be in compliance with them. Nonetheless, the testing failed to reveal a toxin which was a member of the Bacillus cereus bacteria group. That’s because standard testing was simply not precise enough. The good news is that a new, much more precise method had already been developed by a company in Switzerland.

Earlier this year, a number of companies were forced to recall potentially dangerous baby formula. The presence of a toxin called cereulide was suspected and later proven to be present in the formula. The recall cost those companies a lot of money and severely damaged their reputation for quality. But how did that happen? Apparently, the EU-mandated Hazard Analysis and Critical Control Point (HACCP) food safety management program does not properly consider the risk of cereluide contamination in fatty or oily food ingredients.
The Bacillus cereus bacteria group
Cereulide is a member of the Bacillus cereus bacteria group. This group has major societal relevance because it unites within one highly related bacterial complex: 1. frequent food poisoning agents, 2. an important biothreat organism, and 3. globally used biocontrol strains.
On the public health side, B. cereus sensu stricto and close relatives are among the most common causes of bacterial foodborne disease, while B. anthracis causes anthrax and remains central to biosecurity concerns. Meanwhile, B. thuringiensis strains are extensively deployed as “biological” insecticides in agriculture, directly supporting food production. In addition, Bacillus-cereus-group species form very resistant spores, which makes them omnipresent and therefore detectable in the majority of all food products.
Food relevant risks of key species of the Bacillus cereus bacteria group
Species
B. cereus s.s.
Typical ecology & traits
Ubiquitous in soil, raw materials, processing environments; mesophilic; many strains carry the mild enterotoxins (Nhe, Hbl, CytK).
Main food‑related risks
Classical diarrheal food poisoning; spores survive cooking, germinate during cooling/nonconform storage; cereulide in starchy foods can cause vomiting and, rarely, severe complications.
Species
B. thuringiensis
Typical ecology & traits
Insect pathogenic; used widely as a biopesticide; very close to B. cereus genomically; two subclades; strains of clade 1 harbor the same enterotoxin genes (Nhe, Hbl, CytK2) and mid level cytotoxicity as B. cereus s.s.
Main food‑related risks
Biopesticide spores can contaminate vegetables and salads; some strains from food show cytotoxicity comparable to outbreak B. cereus, so residues may occasionally cause B. cereus like illness.
Species
cytotoxicus
Typical ecology & traits
Thermotolerant species, able to grow up to about 50–52 °C; produces highly cytotoxic enterotoxin CytK 1; associated with vegetable dishes and dehydrated plant products.
Main food‑related risks
Linked to severe and sometimes fatal diarrheal outbreaks; heat tolerant spores and growth at elevated temperatures challenge standard heat treatments for ready meals and soups.
Species
B. paranthracis
Typical ecology & traits
Recently described species closely related to B. cereus and B. anthracis; the species harbors all currently characterized chromosomally related, cereulide producing (ces positive) emetic strains
Main food‑related risks
Cereulide producing strains can trigger abrupt, profuse vomiting within hours of ingesting highly contaminated, temperature abused foods. In cases with very high toxin loads, cereulide’s mitochondrial toxicity has been linked to acute liver failure, kidney injury, and rare fatal outcomes in otherwise healthy individuals.
Species
B. anthracis
Typical ecology & traits
Zoonotic pathogen causing anthrax; typically associated with infected animals and their products; spores extremely resilient in the environment.
Main food‑related risks
Foodborne anthrax is rare and usually linked to heavily contaminated meat or traditional products; risk in modern food chains is considered very low but requires clear distinction from other group members when suspected.
Traditional testing
Most routine food laboratories see the Bacillus-cereus-group every day, but current standard methods of testing usually stop at a blurred, group level identification, even though the underlying species range from typical food poisoning strains to biopesticide residues and rare high risk lineages. Quantitative criteria (colony forming units: CFU) for the B. cereus group in ready to eat (RTE) foods typically classify <10³ cfu/g as acceptable, ≈10³–10⁴ cfu/g as borderline, and ≥10⁴–10⁵ cfu/g as unsatisfactory or potentially hazardous, based on assumed cell densities required for toxinogenesis (1).
Unfortunately, these limits are scientifically unsatisfactory because they neither discriminate emetic (ces positive) from non emetic strains nor account for strain specific enterotoxin profiles and matrix /temperature dependent toxin production kinetics. That means that the actual hazard is only weakly predicted by total CFU alone.
Improvements have been available, but…
Whole genome sequencing is essentially the only method that can, in one step, unambiguously place Bacillus cereus group isolates into genomospecies resolved clonal relationships, and comprehensively inventory virulence and resistance genes: no combination of phenotypic tests or single/low plex PCRs can fully achieve that. However, that approach is still not widely used in routine food and clinical laboratories for the simple reason that it requires substantial capital investment, bioinformatics infrastructure and expertise, standardized pipelines, and added turnaround time and cost.
Meanwhile, standard Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry (MALDI TOF MS) – widely used in routine microbiology – speeds up the identification process but uses pattern based matching against clinical centric libraries that only partially cover the B. cereus group.
The background to discovery
Mabritec AG is a company located in the Basel area of Switzerland, and specializing in the identification of microorganisms and the characterization of biological systems using MALDI-TOF mass spectrometry. It currently offers the largest MALDI-TOF database for bacteria in the world – MabritecCentral (2) – and it was during the building of this database a few years ago that something else began to develop.
Using a new MALDI-8020 (among others), Mabritec discovered that separating the Bacillus cereus group based on ribosomal protein masses using MALDI-TOF MS In-Silico is possible. Genome-guided MALDI-TOF MS had not previously been thought possible using commercially available applications for bacterial identification. Now, however, it was.
Diving deep into the concept of genome-guided MALDI-TOF MS
Here is a look into the technical aspects of developing this new method for clearly identifying and quantifying Bacillus cereus group presence in samples.
The MabritecCentral database contains in silico predicted masses of mainly ribosomal proteins from > 440,000 public WGS datasets, curated taxonomically and validated against experimental MALDI TOF spectra. Mabritec identified 3,024 Bacillus cereus group genomes covering all 28 currently known species. The toxin genes (Nhe, Hbl, CytK, EntFM, ces and the virulence plasmids pXO1 and pXO2) were extracted for all strains. As described in the literature, a strong correlation between phylogenetic species and toxin content could thereby be confirmed. This is especially pronounced for the emetic toxin cereulide, exclusively found in B. paranthracis strains. Interestingly, even isolates from cluster 2 of B. thuringiensis are carriers of the mild toxins. Also, the anthrax-causing virulence plasmids are exclusively found in a subcluster of B. anthracis. CytK1, the highly cytotoxic variant of cytotoxin K, is only found in B. cytotoxicus.
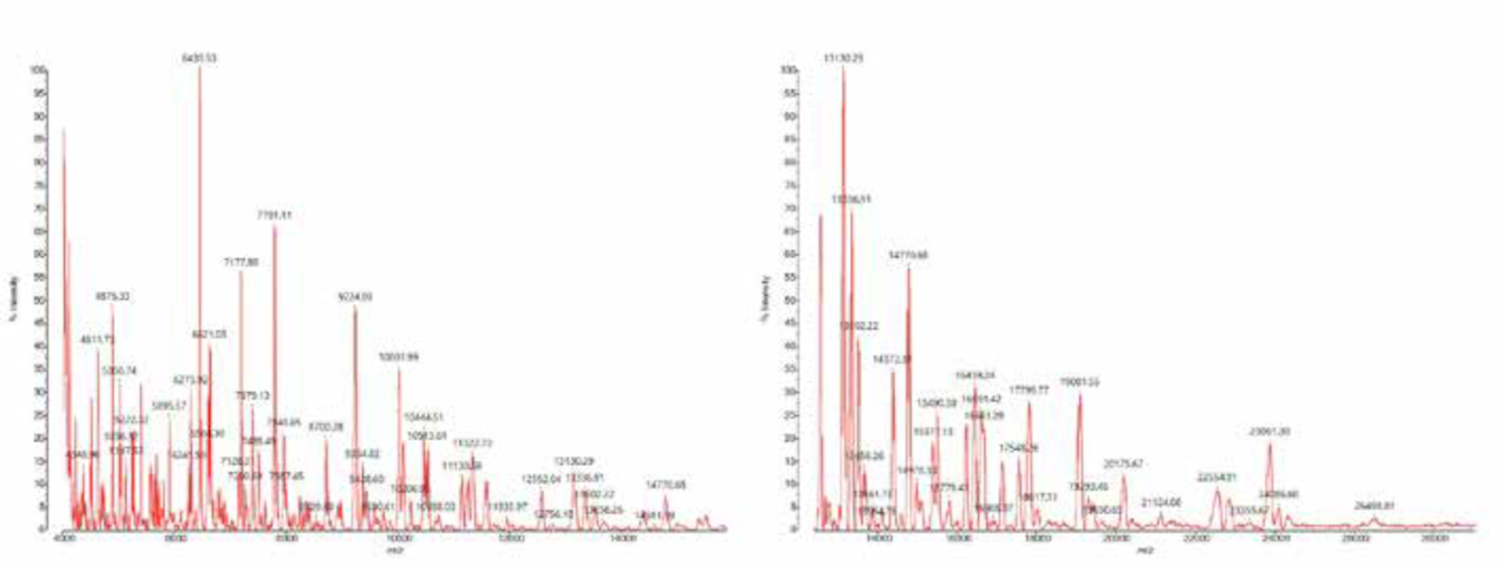
In silico analysis of 32 ribosomal subunit proteins in the 4–30 kDa mass range showed that all species formed distinct clusters that are potentially distinguishable by MALDI-TOF MS. From the in silico prediction, important discriminatory ribosomal proteins were identified in the 15–25 kDa mass range, which are barely detectable using standard smear preparation and CHCA matrix. By using sinapic acid as the matrix together with an adapted acquisition method (adjusted laser power, detector voltage, and peak picking), ribosomal proteins in the 4–28 kDa mass range can be detected (Figure 1).
A further challenge in developing the method was controlling spore formation in order to maintain high spectral quality. Spore formation is known to adversely affect spectral quality because small acid-soluble proteins (SASP) from the spore core can dominate the spectrum and suppress ribosomal marker peaks, particularly in the higher mass range. These limitations could be addressed by two measures. First, a short incubation period of 4–6 hours was applied to generate vegetative cells while keeping spore formation to a minimum. Second, a sample preparation protocol without acid extraction was established, incorporating physical disruption of cells with 0.1 mm glass beads to minimize spore detection. These adjustments allow to high-quality spectra of B. cereus group strains.
This new B. cereus group module was validated with 124 genome sequenced reference strains covering all currently known species. All analyses were performed on an AXIMA Confidence and an 8030 EasyCare MALDI TOF MS. We reached a sensitivity of 98.4% and a specificity of 99.9%. In addition, we could show (in the framework of a small study with 16 clinical isolates) the correct identification of all strains. In addition to the species identification, for 5 strains identified as B. anthracis, we predicted the isolates fall into a B. anthracis ribosomal sublineage profile not harboring the virulence plasmids pXO1 and pXO2. This prediction was confirmed by whole genome sequencing (3).
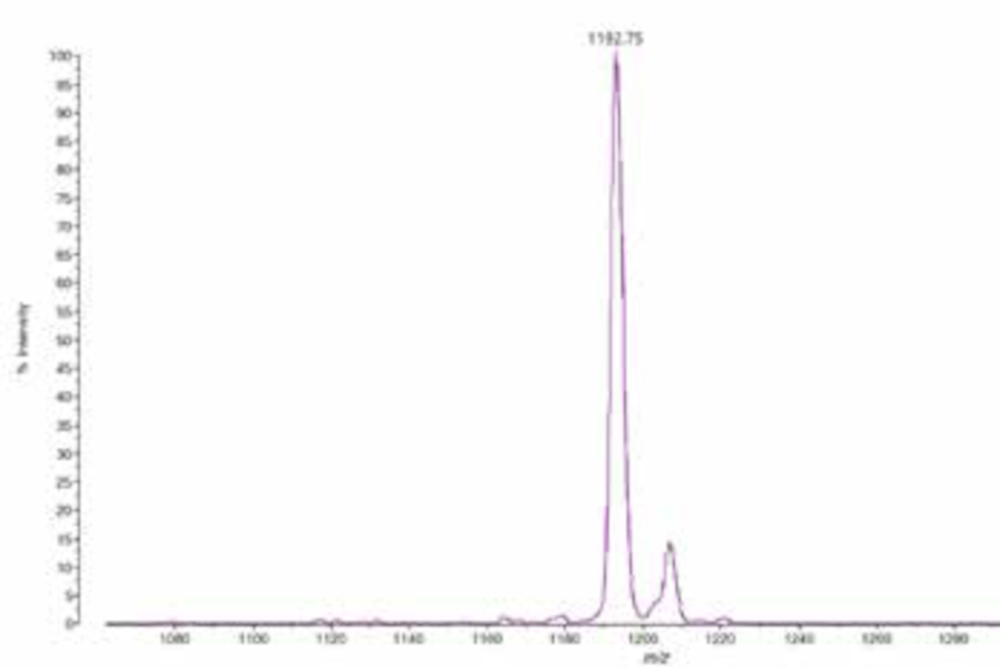
On top of the species level identification of all Bacillus-cereus-group species, we wanted to investigate a potential direct detection of Bacillus-cereus-group toxins by MALDI-TOF MS. We could establish a direct detection of the emetic toxin cereulide directly from cultures or food products by MALDI-TOF MS (4). We found that B. paranthracis strains producing the emetic toxin can be analyzed by directly smearing a small amount of culture material onto a MALDI target and running it in LDI mode without matrix addition, within the mass range of 700–1500 Da. The cereulide is detected at a m/z 1191[M + K]+ (Figure 2). This rapid and straightforward assay represents an efficient and reliable approach to confirm cereulide secretion in suspected toxin-producing B. cereus group strains.
We further investigated the feasibility of detecting cereulide directly in food samples. For the cereulide test, 1 g of homogenized food (e.g. rice, milk rice, ravioli, mashed potatoes, etc.) is mixed with 1 mL of 75% (v/v) ethanol in a 2 mL tube, vortexed 1 min, then centrifuged at 6,000 × g for 10 min. After centrifugation, 1 μL of the clear supernatant is spotted in duplicate onto the stainless steel target, air dried at room temperature and overlayed with 1 μL CHCA matrix. Cereulide is highly hydrophobic and strongly associates with food components, so extraction efficiency is dependent on the type of food analyzed. Recovery from starchy foods (e.g. rice) is usually higher than from fatty, protein-rich, or complex composite dishes where cereulide partitions into lipid phases or binds to proteins. The limits of detection can vary in a range from 30 ng/mL to 30pg. For every sample tested, we run a dilution series of cereulide spikes for LOD definition. In practice, for risk assessment one may assume that any cereulide level capable of causing typical emetic food poisoning (reported in the low µg/g range in outbreaks) will be clearly detectable by this LDI workflow in responsive matrices.


When science benefits consumers, laboratories and companies
Combining genome-based references for identifying toxin producers with rapid and cost-effective MALDI-TOF MS analysis – in contrast to expensive genome sequencing – has now resulted in a new and advantageous ability to routinely differentiate all Bacillus cereus group species and their potential virulence in food. For the first time, all species can be identified under routine conditions using MALDI-TOF MS and assigned a toxin profile using a genome-based database. Depending on the findings and the matrix/product, critical food products can thus be proactively removed from circulation.
This new method can be used in the food, pharmaceutical and cosmetics industries along the entire production chain, from raw materials through production/processing to the final product. It allows quality managers to better interpret the frequently detected Bacillus cereus group, improve food safety, and better protect consumers. The method is already routinely used in the Mabritec laboratory where more than 1,600 Bacillus cereus group isolates originating from food and industrial samples have been identified.
Good tools help good people make things better
Genome guided MALDI TOF MS enables reliable species level resolution within the Bacillus cereus group and links isolates to characteristic toxin and virulence profiles that remain invisible to conventional CFU based and routine identification approaches. The work of the scientists at Mabritec demonstrates that rapid MALDI/LDI based detection of cereulide from cultures and suitable food matrices is technically feasible, sufficiently sensitive, and compatible with routine workflows, thereby strengthening risk assessment and surveillance for emetic B. cereus group strains. Using Shimadzu equipment, a clever Swiss company came up with a new way to better protect the health of consumers, and which also saves companies from costly and embarrassing product recalls.
1 Carroll LM, Wiedmann M, Kovac J. Proposal of a Taxonomic Nomenclature for the Bacillus cereus Group Which Reconciles Genomic Definitions of Bacterial Species with Clinical and Industrial Phenotypes. mBio. 2020 Feb 25;11(1):e00034-20. doi: 10.1128/mBio.00034-20. PMID: 32098810; PMCID: PMC7042689.
2 Institute of Medicine and National Research Council. 2003. Scientific Criteria to Ensure Safe Food. Washington, DC: The National Academies Press.
3 www.mabriteccentral.com
4 Muigg V, Cuénod A, Purushothaman S, Siegemund M, Wittwer M, Pflüger V, Schmidt KM, Weisser M, Ritz N, Widmer A, Goldenberger D, Hinic V, Roloff T, Søgaard KK, Egli A, Seth-Smith HMB. Diagnostic challenges within the Bacillus cereus-group: finding the beast without teeth. New Microbes New Infect. 2022 Oct 26;49-50:101040. doi: 10.1016/j.nmni.2022.101040. PMID: 36385748; PMCID: PMC9641004.
5 Ducrest PJ, Pfammatter S, Stephan D, Vogel G, Thibault P, Schnyder B. Rapid detection of Bacillus ionophore cereulide in food products. Sci Rep. 2019 Apr 9;9(1):5814. doi: 10.1038/s41598-019-42167-0. PMID: 30967595; PMCID: PMC6456620.